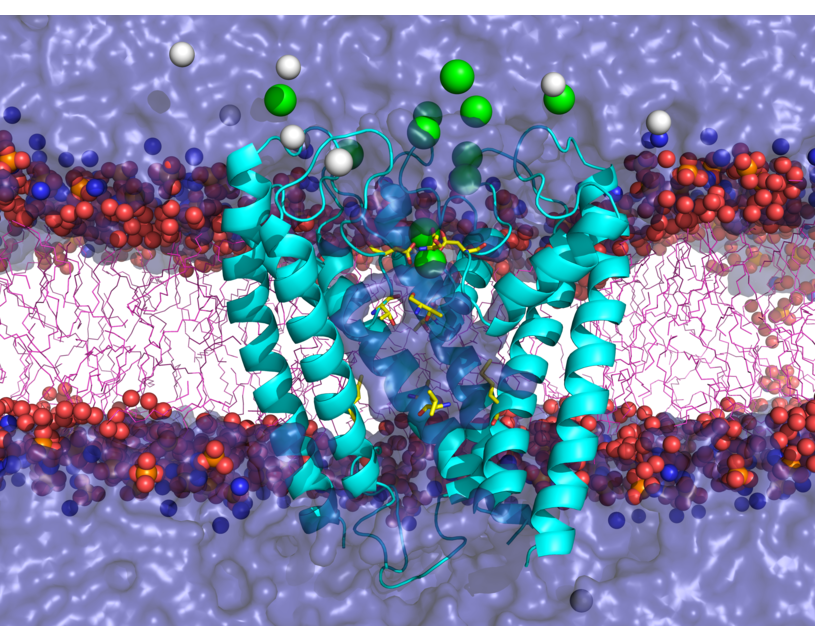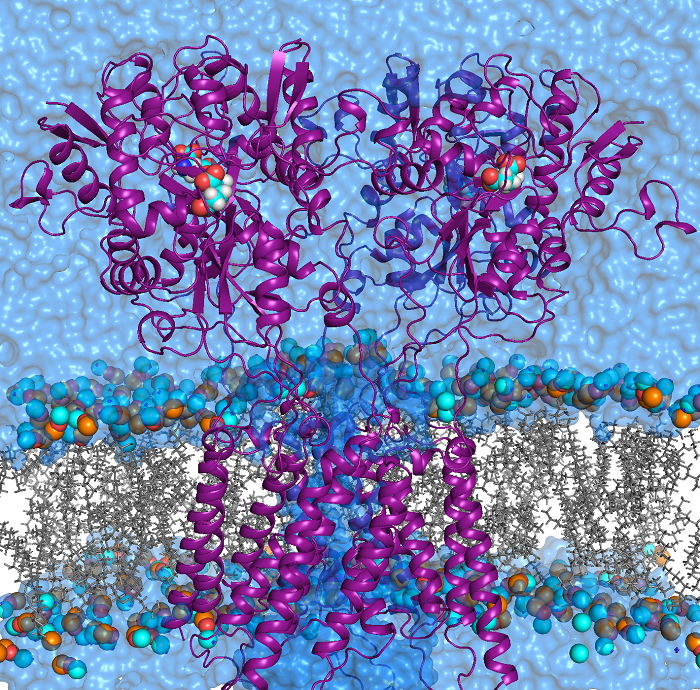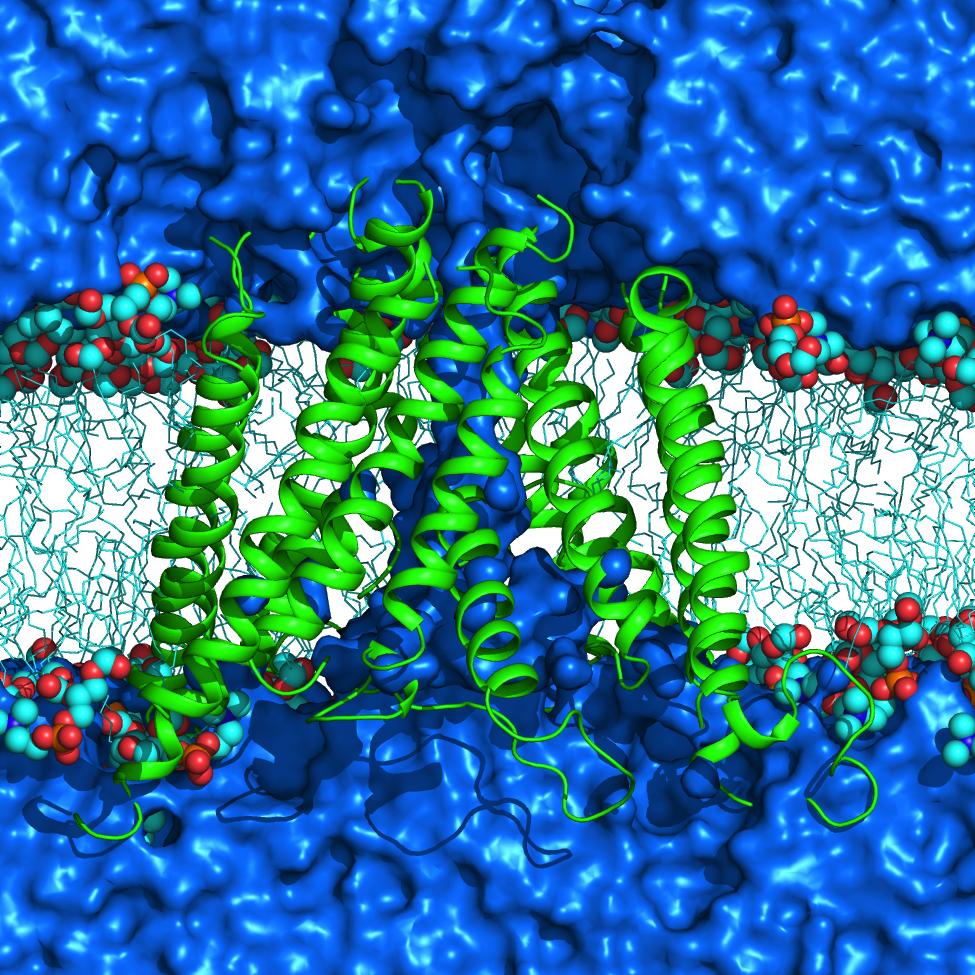
NMDA receptors (NMDARs) are glutamate- and glycine-gated ionotropic glutamate receptors (iGluRs) found mainly in postsynaptic membranes in the central nervous system. NMDARs are required for many forms of synaptic plasticity, including long-term potentiation and long-term depression, physiological processes that are involved in learning and memory. Dysfunction of NMDARs has been associated with multiple nervous system disorders. We investigate a variety of properties in the transmembrane domain (TMD) of NMDARs.
One area of interest includes study of ion selectivity in NMDARs. NMDAR channels are permeable to monovalent cations including Na+ and K+, as well as Ca2+, and are blocked by Mg2+ in a membrane potential-dependent manner. Using molecular dynamics simulations and free energy calculations, we are investigating the basis for Ca2+ permeation versus Mg2+ block in NMDARs. This work led to proposing a novel reaction coordinate (RC) for the calculation of the potential of mean force (PMF) inside narrow channels. This new RC takes into account the slowest degree of freedom for ion permeation reaction, a property lacking when using a straightforward Z-coordinate, a common RC used for ion permeation PMF calculation inside ion channels. Currently, we are exploring the full configurational space of the Mg2+ and Ca2+ in NMDAR channel to accomplish an accurate PMF for describing permeation versus block of divalent ions.
Another area of research, in collaboration with Jon W. Johnson's group, is interaction of several drugs with NMDARs. NMDARs can be inhibited in several ways, one of which is open channel block, in which molecules bind in the pore of NMDARs and block ion flow. Memantine (Mem), a drug used to treat Alzheimer’s disease, is an NMDAR open channel blocker that binds in the NMDAR pore. We study the interaction of Mem with NMDARs and investigate its permeation pathway to the binding site.
Within the M2 region of each NMDAR subunit lies a remarkably conserved tryptophan (W) that can be found at the homologous site in a variety of channels and receptors in phylogenetically distant organisms. In NMDARs, this highly conserved W is located at the interface between the M2 region and the adjacent M3 region. The conserved W plays a role in NMDAR function; mutation of the conserved W in the GluN2B subunit (W607) severely diminishes Mg2+ block. However, mutating the homologous residue in GluN1 subunits (W608) does not affect Mg2+ block, but alters desensitization. To further explore the function of the conserved W in NMDARs, in collaboration with Jon W. Johnson's group, we examined GluN1/GluN2A receptors with mutations at the conserved W and surrounding residues. Our work provided molecular-detailed understanding of altered function of NMDAR seen in electrophysiological experiments by our collaborators.
Recent Publications:
S. Mesbahi-Vasey, J. W. Johnson, M. G. Kurnikova, “Reaction coordinate for an ion association with a protein binding site: configurations of the ligand coordination shell”, submitted.
S. Mesbahi-Vasey, L. Veras, M. Yonkunas, J. W. Johnson, M. G. Kurnikova, “All atom NMDA receptor transmembrane domain model development and simulations in lipid bilayers and water”, PLoS ONE 12(6): e0177686 (2017).